RESEARCH
PLANT BIODIVERSITY: SYSTEMATICS, PHYLOGENETICS AND EVOLUTION
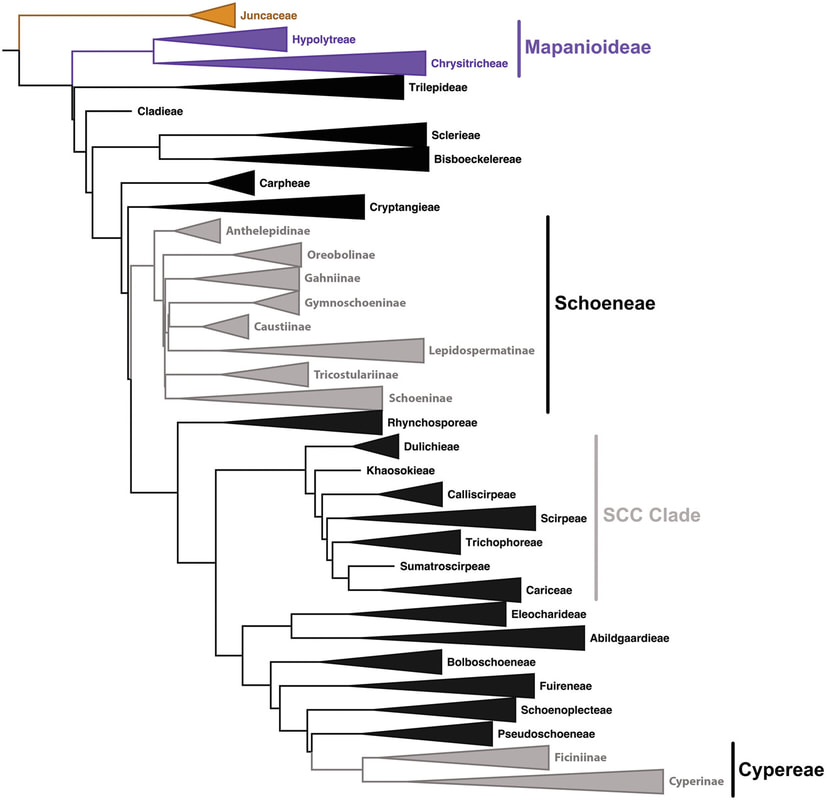
Our research interest is mainly focused in the study of biodiversity and diversification and evolution of flowering plants at different evolutionary scales. My aims are (i) to quantity the biodiversity (How many species are there? How are those species delimited and related?) and (ii) to disentangle the mechanisms that generate and maintain such biodiversity. We have worked with several lineages from different flowering plant families (Gentianaceae: Centaurium or Linaceae: Linum) but we have mainly used as study-group the family Cyperaceae and especially the genus Carex.
Our research interest is mainly focused in the study of biodiversity and diversification and evolution of flowering plants at different evolutionary scales. My aims are (i) to quantity the biodiversity (How many species are there? How are those species delimited and related?) and (ii) to disentangle the mechanisms that generate and maintain such biodiversity. We have worked with several lineages from different flowering plant families (Gentianaceae: Centaurium or Linaceae: Linum) but we have mainly used as study-group the family Cyperaceae and especially the genus Carex.
Phylogenetics, Systematics and Taxonomy
As mention before, one of the main goals of my research is to answer the questions: How many species are there? How are those species delimited and related?
We have used many different molecular techniques (NGS like GBS or RADseq or HybSeq, Sanger sequencing DNA, AFLP, SSR, etc.) depending on the evolutionary scale of the study to infer phylogenetic hypotheses.
As mention before, one of the main goals of my research is to answer the questions: How many species are there? How are those species delimited and related?
We have used many different molecular techniques (NGS like GBS or RADseq or HybSeq, Sanger sequencing DNA, AFLP, SSR, etc.) depending on the evolutionary scale of the study to infer phylogenetic hypotheses.
Inferring evolutionary patterns by using phylogenetic comparative methods
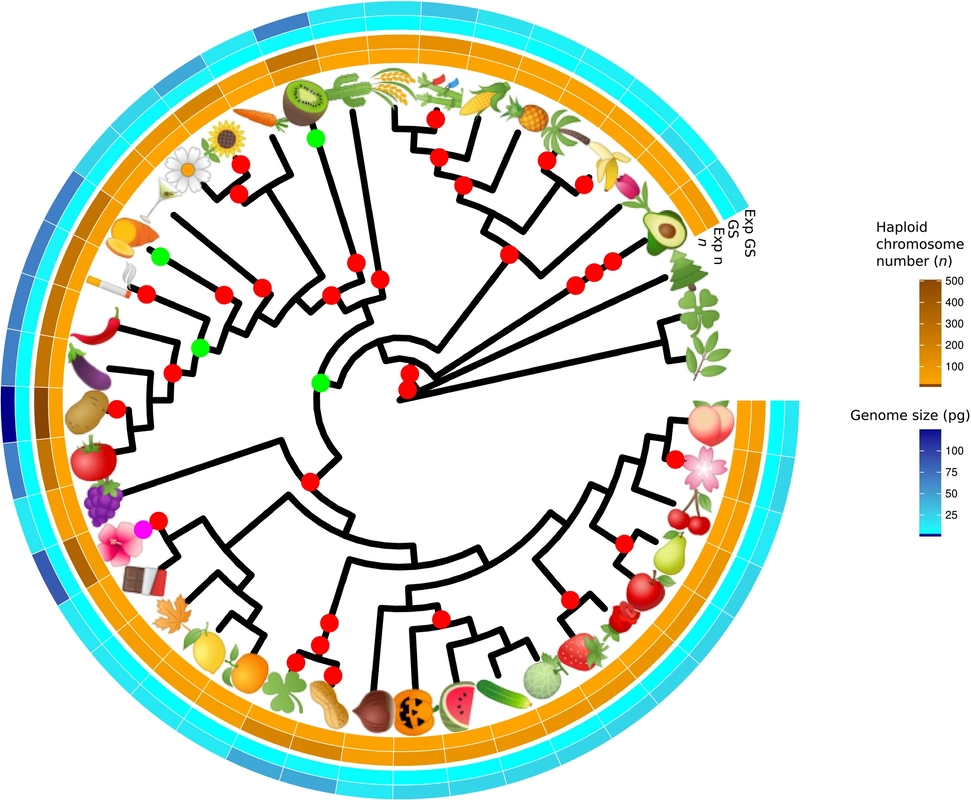
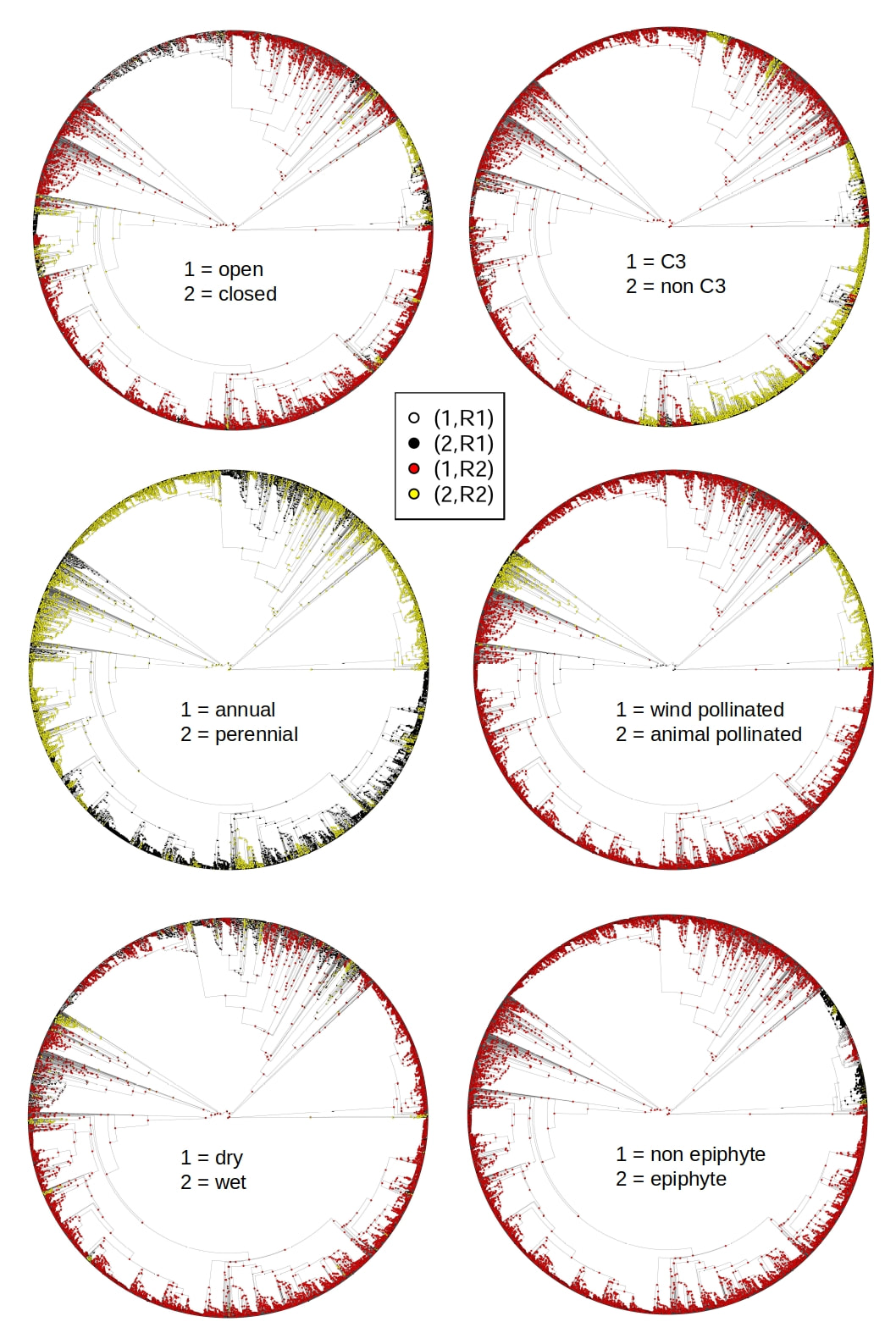
Trait modelling and reconstructions of ancestral character states make possible to know when, how and why traits evolve. Phylogenetic comparative methods use information on the evolutionary relationships of organisms (phylogenetic trees) to compare species (or populations). The most common applications are to test for correlated evolutionary changes in two or more traits, or to determine whether a trait contains a phylogenetic signal, the tendency for related species to resemble each other.
Trait modelling and reconstructions of ancestral character states make possible to know when, how and why traits evolve. Phylogenetic comparative methods use information on the evolutionary relationships of organisms (phylogenetic trees) to compare species (or populations). The most common applications are to test for correlated evolutionary changes in two or more traits, or to determine whether a trait contains a phylogenetic signal, the tendency for related species to resemble each other.
Chromosome evolution
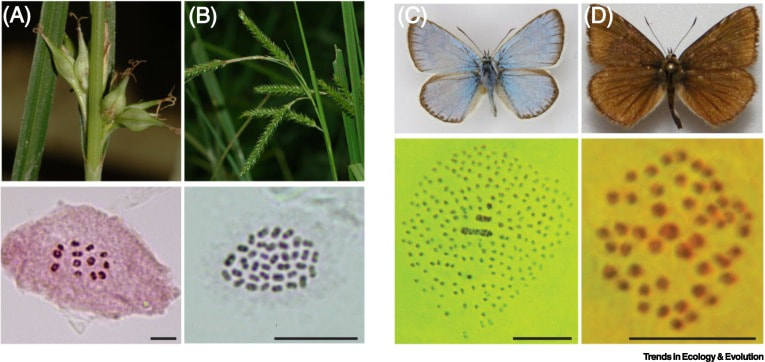
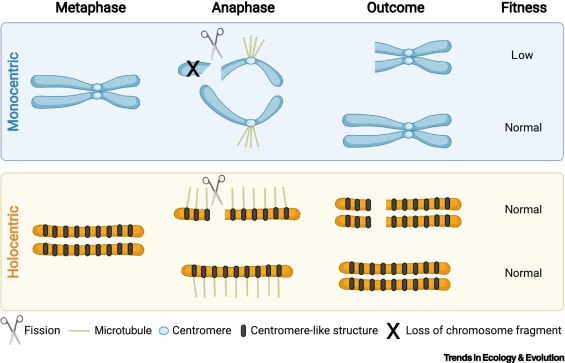
This is one of the most peculiar aspects in my research since we study chromosome evolution in holocentric organisms using phylogenetic comparative methods. Holocentric chromosomes—chromosomes with diffuse rather than solitary, localized centromeres—have been reported from sedges and rushes (Cyperaceae and Juncaceae, which together form a clade) and five other angiosperm clades, as well as scattered clades in the algae, bryophytes, Rhizaria, nematodes, velvet worms, and arthropods. Diffuse centromeres allow rapid evolution of chromosome rearrangements via fission (agmatoploidy), fusion (simploidy), and translocations. Consequently, Carex exhibits a nearly continuous range of chromosome numbers (2n = 12 to 124) and substantial chromosome variation within many species. Diffuse centromere has the potential to reduce or eliminate the underdominance of chromosome rearrangements, allowing them to become fixed at a higher rate than in organisms with monocentric chromosomes. Holocentric chromosome evolution (i) promotes genetic differentiation and (ii) seems to be selected by the climate niche, both at micro and macroevolutionary scales.
This is one of the most peculiar aspects in my research since we study chromosome evolution in holocentric organisms using phylogenetic comparative methods. Holocentric chromosomes—chromosomes with diffuse rather than solitary, localized centromeres—have been reported from sedges and rushes (Cyperaceae and Juncaceae, which together form a clade) and five other angiosperm clades, as well as scattered clades in the algae, bryophytes, Rhizaria, nematodes, velvet worms, and arthropods. Diffuse centromeres allow rapid evolution of chromosome rearrangements via fission (agmatoploidy), fusion (simploidy), and translocations. Consequently, Carex exhibits a nearly continuous range of chromosome numbers (2n = 12 to 124) and substantial chromosome variation within many species. Diffuse centromere has the potential to reduce or eliminate the underdominance of chromosome rearrangements, allowing them to become fixed at a higher rate than in organisms with monocentric chromosomes. Holocentric chromosome evolution (i) promotes genetic differentiation and (ii) seems to be selected by the climate niche, both at micro and macroevolutionary scales.
Biogeography and phylogeography
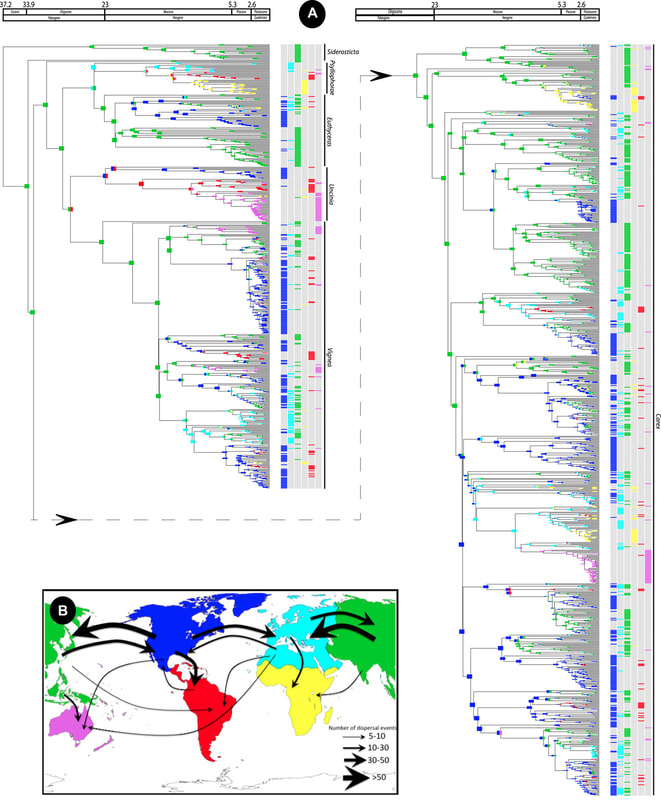
We have studied intensely biogeography of flowering plants using the last phylogenetic methods to estimate times of diversification and reconstruction of ancestral areas in different linages of plants.
For example, we have inferred for genus Carex, or favourite study system, that has a special ability to colonize novel and far environments via long-distance dispersal. As result, many long-distance dispersal events have been inferred. Léon Croizat (Manual of Phytogeography, 1952) once said that you could teach an entire course on phytogeography using only examples from Carex. There are sedge species and groups with almost every imaginable biogeographic pattern. The question is which processes have created such extraordinary and extreme distributions?
We have studied intensely biogeography of flowering plants using the last phylogenetic methods to estimate times of diversification and reconstruction of ancestral areas in different linages of plants.
For example, we have inferred for genus Carex, or favourite study system, that has a special ability to colonize novel and far environments via long-distance dispersal. As result, many long-distance dispersal events have been inferred. Léon Croizat (Manual of Phytogeography, 1952) once said that you could teach an entire course on phytogeography using only examples from Carex. There are sedge species and groups with almost every imaginable biogeographic pattern. The question is which processes have created such extraordinary and extreme distributions?